The Science Of Scientific Writing Set 5 Set 5-Essays • Second page • Third page • Ordering ideas • Exercise 1 • Signposting• Final.
OVERVIEW: The way to well-written science
PART I: Paragraphs and Sentences...
SET 1: The Parts of Arguments
SET 2: Indicator Words
SET 3: Refining Claims
SET 4: Locating Arguments in Prose
SET 5: Rationale's Essay Planner
SET 6: Assessing
SET 7 : More on Assessing
Maps provide clues that must also be present in our texts
When we convert an argument map to text form we need to make extra efforts to ensure that the relationships between our ideas remain as obvious as they are on the map.
In the generic map below, even in the total absence of any knowledge of the map's content, a great deal of information is provided by the structure of the map itself:
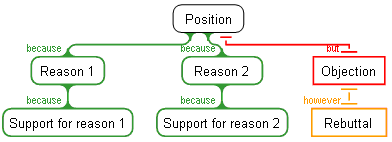
The viewer immediately knows:
- The position being argued
- The total number of primary lines of argument, and how many of these are SUPPORTING, or COUNTERING the position.
- The degree of branching (both vertical and horizontal) for each primary argument line.
This bird's eye view of the argument's architecture tells us a lot, and gives us a head start when we do start to deal with the argument's content.
From map to text
If you were going to write an essay based on this map, one important job would be to work out the best way to arrange the ideas on the page.
The map is very clear about how the reasoning goes. We can keep this clarity by carefully arranging the claims in the three parts of an essay:
-
The introduction clearly identifies the position, in a position-early essay, or the question-addressed-by-the-position (we will call this the issue), in a position-final essay. The introduction will usually contain information that is NOT present in the map itself.
-
The body examines the branches of the argument. The essay steps the reader through the argument one branch at a time, working from the top of the branch to the bottom. So our essay could have this form:
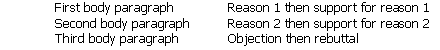
- The conclusion reminds the reader of the position, in a position-early essay, or announces it for the first time, in a position-final essay.
Navigational Strategies of Scientfic Papers
A scientific paper is just an expanded essay, in which the results that form the main evidential basis of the argument/s in the Discussion are given particular emphasis by being treated in a separate section. The ideas that apply to essay organisation also apply to writing a scientific paper.
How can we achieve such navigational transparency in a (multiparagraph) written piece?
"External" navigational strategies in written text: signposting
I will first focus on the various forms of signposting used in scientific papers, that involve text (or diagrams) that are found outside of the paragraphs of the main text. Explicit strategies such as theese will not be our emphasis in this course, but I include them here because they highlight the importance of navigational assistance. Scientific papers present a good model system for studying the evolution of navigational devices, because the volume and complexity of material is probably higher than in any other area.
Here are some of the navigational features of scientific papers:
- The main body of the paper has (most typically) four functionally different main sections (Introduction/Materials & Methods/Results/Discussion)
- Within a section, sub-sections may be signalled by the use of headings
- Diagrams and flow charts are often introduced to guide the reader
- The whole paper is summarised in an abstract
- The abstract may itself be broken up into functional subdivisions! (see example below: the paragraphing of the abstract is also a feature of this and some other journals)
Possible
Prediction of Chemoradiosensitivity of Esophageal Cancer by Serum Protein Profiling
Yasuharu Hayashida et al.(2005) Clinical Cancer Research, 11: 8042-47.
Abstract
Purpose: Establishment of a reliable method of predicting the efficacy of chemotherapy and radiotherapy is necessary to provide the most suitable treatment for each cancer patient. We investigated whether proteomic profiles of serum samples obtained from untreated patients were capable of being used to predict the efficacy of combined preoperative chemoradiotherapy against esophageal cancer.
Experimental Design: Proteomic spectra were obtained from a training set of 27 serum samples (15 pathologically diagnosed responders to preoperative chemoradiotherapy and 12 nonresponders) by surface-enhanced laser desorption and ionization coupled with hybrid quadrupole time-of-flight mass spectrometry. A proteomic pattern prediction model was constructed from the training set by machine learning algorithms, and it was then tested with an independent validation set consisting of serum samples from 15 esophageal cancer patients in a blinded manner.
Results: We selected a set of four mass peaks, at 7,420, 9,112, 17,123, and 12,867 m/z, from a total of 859 protein peaks, as perfectly distinguishing responders from nonresponders in the training set with a support vector machine algorithm. This set of peaks (i.e., the classifier) correctly diagnosed chemoradiosensitivity in 93.3% (14 of 15) of the cases in the validation set.
Conclusions: Recent mass spectrometric approaches have revealed that serum contains a large volume of information that reflects the microenvironment of diseased organs. Although a multi-institutional large-scale study will be necessary to confirm each component of the classifier, there is a subtle but definite difference in serum proteomic profile between responders and nonresponders to chemoradiotherapy.
All of these (mainly) structural features provide the reader, well before they have confronted the content, with generic navigational assistance analogous to that provided visually by an argument map.
No doubt, as science becomes even more voluminous and complex, and the move away from hardcopy-based publishing accelerates, the evolution of features that provide even greater transparency will continue. Already we find that data that would have originally been included in a paper (e.g. gene sequences) now go directly into online databases (e.g. GenBank), greatly facilitating their examination. Online publishing also offers the possibility of hypertext linking, which can allow the easy separation of material using any number of criteria (importance to the text overall; historical background; methodological interest; intellectual complexity, etc.).
Content of this page drawn in whole or part from the Austhink Rationale Exercises with permission from Austhink.